-Search query
-Search result
Showing 1 - 50 of 59 items for (author: ma & xm)

EMDB-35042: 
Cryo-EM structure of the the 2-oxoglutarate dehydrogenase (E1) with TCAIM complex
Method: single particle / : Yu X, Yang W

EMDB-17766: 
CryoEM structure of Nal1 protein, allele SPIKE, from Oryza sativa japonica group
Method: single particle / : Huang LY, Rety S, Xi XG

EMDB-17768: 
CryoEM structure of Nal1 protein, allele IR64, from Oryza sativa indica cultivar
Method: single particle / : Huang LY, Rety S, Xi XG
EMDB-35297: 
Structural insights into human brain gut peptide cholecystokinin receptors
Method: single particle / : Ding Y, Zhang H, Liao Y, Chen L, Ji S

EMDB-15417: 
human MutSalpha (MSH2/MSH6) binding to DNA with a GT mismatch
Method: single particle / : Bruekner SR, Sixma TK

EMDB-15519: 
human MutSalpha (MSH2/MSH6) on DNA containing a GT mismatch in the presence of ADP
Method: single particle / : Bruekner SR, Liaci AM, Sixma TK

PDB-8ag6: 
human MutSalpha (MSH2/MSH6) binding to DNA with a GT mismatch
Method: single particle / : Bruekner SR, Sixma TK

EMDB-15518: 
human MutSalpha mutant (MSH2-V63E/MSH6) on DNA containing a GT mismatch
Method: single particle / : Bruekner SR, Sixma TK

EMDB-26030: 
Structure of G6PD-WT tetramer with no symmetry imposed
Method: single particle / : Wei X, Marmorstein R

EMDB-26031: 
Structure of G6PD-WT dimer with no symmetry applied
Method: single particle / : Wei X, Marmorstein R

EMDB-26428: 
Structure of G6PD-D200N tetramer bound to NADP+ and G6P with no symmetry applied
Method: single particle / : Wei X, Marmorstein R

EMDB-26442: 
Structure of G6PD-D200N tetramer bound to NADP+ with no symmetry applied
Method: single particle / : Wei X, Marmorstein R

PDB-7toe: 
Structure of G6PD-WT tetramer with no symmetry imposed
Method: single particle / : Wei X, Marmorstein R

PDB-7tof: 
Structure of G6PD-WT dimer with no symmetry applied
Method: single particle / : Wei X, Marmorstein R

PDB-7ual: 
Structure of G6PD-D200N tetramer bound to NADP+ and G6P with no symmetry applied
Method: single particle / : Wei X, Marmorstein R

PDB-7uc2: 
Structure of G6PD-D200N tetramer bound to NADP+ with no symmetry applied
Method: single particle / : Wei X, Marmorstein R

EMDB-33359: 
Structural insights into human brain gut peptide cholecystokinin receptors
Method: single particle / : Ding Y, Zhang H, Liao Y, Chen L, Ji S

EMDB-33360: 
Structural insights into human brain gut peptide cholecystokinin receptors
Method: single particle / : Ding Y, Zhang H, Liao Y, Chen L, Ji S

EMDB-33361: 
Structural insights into human brain gut peptide cholecystokinin receptors
Method: single particle / : Ding Y, Zhang H, Liao Y, Chen L, Ji S

EMDB-25224: 
Structure of G6PD-WT dimer
Method: single particle / : Wei X, Marmorstein R

EMDB-25225: 
structure of G6PD-WT tetramer
Method: single particle / : Wei X, Marmorstein R

EMDB-25226: 
Structure of G6PD-D200N tetramer bound to NADP+
Method: single particle / : Wei X, Marmorstein R
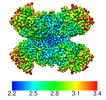

EMDB-25227: 
Structure of G6PD-D200N tetramer bound to NADP+ and G6P
Method: single particle / : Wei X, Marmorstein R

PDB-7snf: 
Structure of G6PD-WT dimer
Method: single particle / : Wei X, Marmorstein R

PDB-7sng: 
structure of G6PD-WT tetramer
Method: single particle / : Wei X, Marmorstein R

PDB-7snh: 
Structure of G6PD-D200N tetramer bound to NADP+
Method: single particle / : Wei X, Marmorstein R

PDB-7sni: 
Structure of G6PD-D200N tetramer bound to NADP+ and G6P
Method: single particle / : Wei X, Marmorstein R

EMDB-32485: 
Cryo-electron microscopic structure of the 2-oxoglutarate dehydrogenase (E1) component of the human alpha-ketoglutarate (2-oxoglutarate) dehydrogenase complex
Method: single particle / : Yu X, Yang W, Zhong YH, Ma XM, Gao YZ

EMDB-33331: 
Cyo-EM model for native cystathionine beta-synthase of Mycobacterium tuberculosis.
Method: single particle / : Bandyopadhyay P, Pramanick I, Biswas R, Sabarinath PS, Sreedharan S, Singh S, Rajmani R, Laxman S, Dutta S, Singh A
EMDB-33348: 
Cryo-EM map of cystathionine beta-synthase of Mycobacterium tuberculosis in the presence of S-adenosylmethionine.
Method: single particle / : Bandyopadhyay P, Pramanick I, Biswas R, Sabarinath PS, Sreedharan S, Singh S, Rajmani R, Laxman S, Dutta S, Singh A
EMDB-33363: 
Cryo-EM map of cystathionine beta-synthase of Mycobacterium tuberculosis in the presence of S-adenosylmethionine and serine.
Method: single particle / : Bandyopadhyay P, Pramanick I, Biswas R, Sabarinath PS, Sreedharan S, Singh S, Rajmani R, Laxman S, Dutta S, Singh A

PDB-7xnz: 
Native cystathionine beta-synthase of Mycobacterium tuberculosis.
Method: single particle / : Bandyopadhyay P, Pramanick I, Biswas R, Sabarinath PS, Sreedharan S, Singh S, Rajmani R, Laxman S, Dutta S, Singh A

PDB-7xoh: 
Cystathionine beta-synthase of Mycobacterium tuberculosis in the presence of S-adenosylmethionine.
Method: single particle / : Bandyopadhyay P, Pramanick I, Biswas R, Sabarinath PS, Sreedharan S, Singh S, Rajmani R, Laxman S, Dutta S, Singh A

PDB-7xoy: 
Cystathionine beta-synthase of Mycobacterium tuberculosis in the presence of S-adenosylmethionine and serine.
Method: single particle / : Bandyopadhyay P, Pramanick I, Biswas R, Sabarinath PS, Sreedharan S, Singh S, Rajmani R, Laxman S, Dutta S, Singh A

EMDB-31246: 
Cryo-electron tomogram of phycobilisome-photosystem II supercomplex on the thylakoid
Method: electron tomography / : Sui SF, Li XM, Li MJ, Ma JF

EMDB-31241: 
Structure of the PBS-PSII supercomplex from red algae
Method: subtomogram averaging / : Sui SF, Li XM, Li MJ, Ma JF

EMDB-31242: 
Structure of the double phycobilisome-photosystem II supercomplex from red alga
Method: subtomogram averaging / : Sui SF, Li XM, Li MJ, Ma JF

EMDB-31243: 
Cryo-electron tomography of phycobilisome-photosystem II supercomplex on the thylakoid
Method: electron tomography / : Sui SF, Li XM, Li MJ, Ma JF

EMDB-31244: 
Cryo-electron tomogram of phycobilisome-photosystem II supercomplex on the thylakoid
Method: electron tomography / : Sui SF, Li XM, Li MJ, Ma JF

EMDB-31245: 
Cryo-electron tomogram of phycobilisome-photosystem II supercomplex on the thylakoid
Method: electron tomography / : Sui SF, Li XM, Li MJ, Ma JF

EMDB-31247: 
Cryo-electron tomogram of phycobilisome-photosystem II supercomplex on the thylakoid
Method: electron tomography / : Sui SF, Li XM, Li MJ, Ma JF

EMDB-30650: 
Cryo-EM structure of the Ams1 and Nbr1 complex
Method: single particle / : Zhang J, Ye K

EMDB-30652: 
Cryo-EM structure of the Ape4 and Nbr1 complex
Method: single particle / : Zhang J, Ye K

EMDB-11791: 
MutS in Scanning state
Method: single particle / : Fernandez-Leiro R, Bhairosing-Kok D

EMDB-11792: 
MutS in mismatch bound state
Method: single particle / : Fernandez-Leiro R, Bhairosing-Kok D

EMDB-11793: 
MutS in Intermediate state
Method: single particle / : Fernandez-Leiro R, Bhairosing-Kok D

EMDB-11794: 
MutS-MutL in clamp state
Method: single particle / : Fernandez-Leiro R, Bhairosing-Kok D

EMDB-11795: 
MutS-MutL in clamp state (kinked clamp domain)
Method: single particle / : Fernandez-Leiro R, Bhairosing-Kok D

PDB-7ai5: 
MutS in Scanning state
Method: single particle / : Fernandez-Leiro R, Bhairosing-Kok D, Sixma TK, Lamers MH

PDB-7ai6: 
MutS in mismatch bound state
Method: single particle / : Fernandez-Leiro R, Bhairosing-Kok D, Sixma TK, Lamers MH
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
